Most people know our genetic code involves the letters A, T, C and G (adenine, thymine, guanine and cytosine) to make up DNA. But what fewer people know about is the code found in the packaging of DNA. And there is a lot of it there.
Until recently, we didn’t have a good way to systematically get at that information. Science took a big step forward in being able to do this with the publication of the most precise map to date of how human cells organize their 6 feet of DNA into their nuclei, a space that has a diameter of only around 0.0000016 inches. This kind of map will not only tell us how the instructions in our DNA lead to making each one of us, but it may also provide new ways to understand and even treat diseases like cancer.
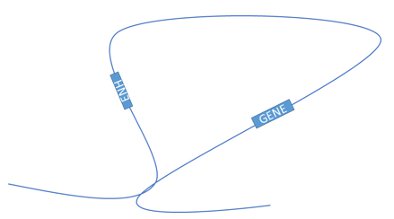
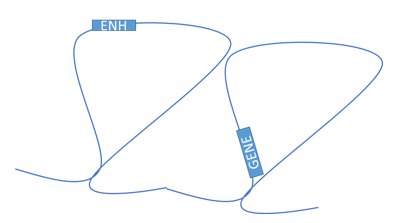
In this study, Baylor College of Medicine researcher Suhas Rao and coworkers focused on a part of nuclear organization called DNA loops. Basically, our DNA forms a set of loops that each act as an independent region. Each loop has a set of controls for genes in that loop that don’t affect genes outside of that loop.
One of the most surprising results of this study was that instead of the one million loops that many scientists expected, the authors found only around 10,000 of them. Not simple to study but definitely a more manageable number than one million!
One way to think about these results is that it is a bit like the original human genome project. Just like that original DNA sequence, it is providing us with a reference to which all other studies like this can be compared.
Scientists will be able to, for example, determine how DNA is packaged in the cells of people with certain diseases and compare it to this reference map. From that comparison they may be able to better understand why a certain DNA change leads to a certain disease.